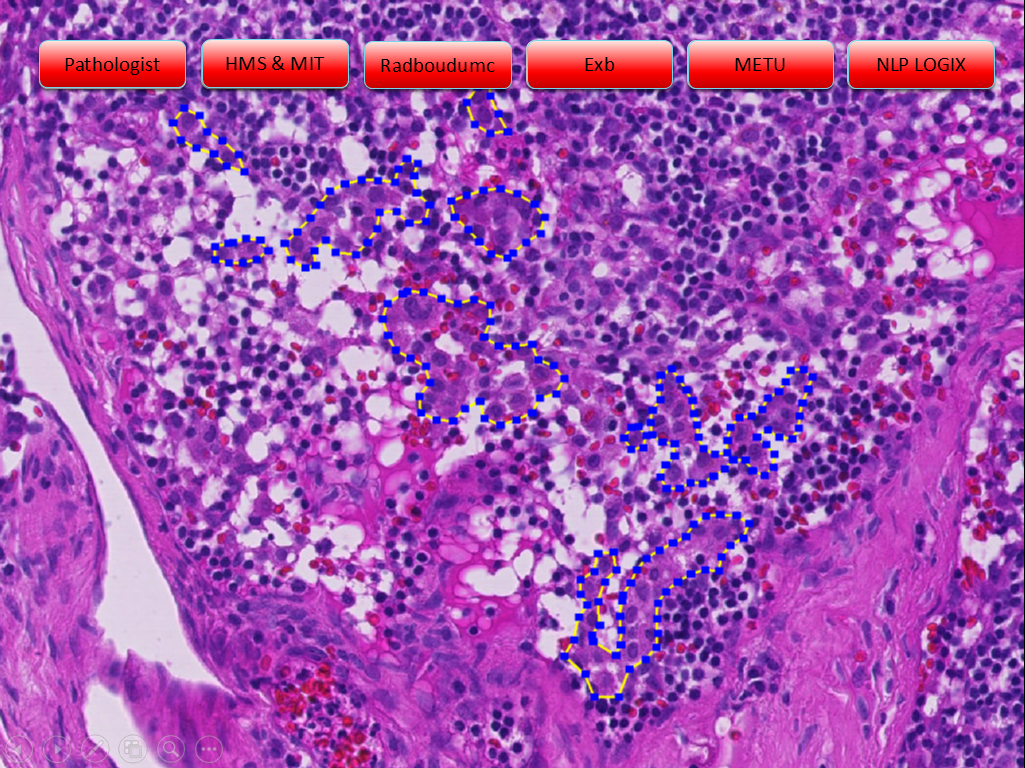
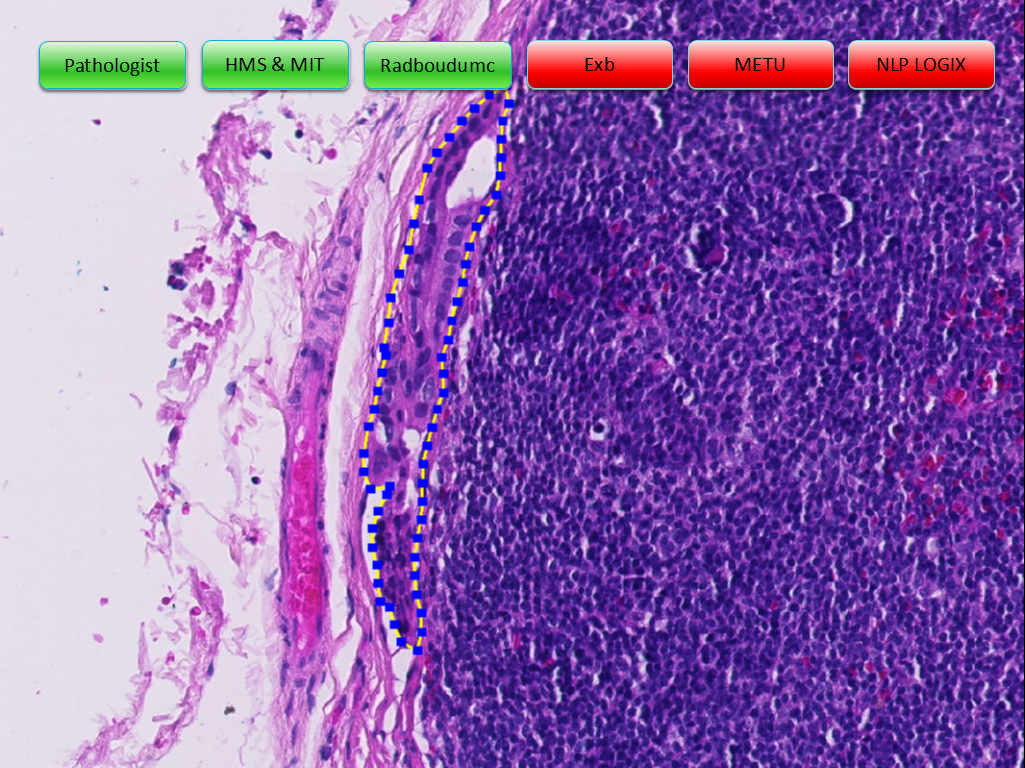
Using the power of social media for professional work isn’t new to pathology. By today, social media channels have become one of the speediest and easiest to use online tools in the disruption of healthcare communication.
Despite an initial reservation, by 2017 pathologist around the world have discovered especially Twitter and Facebook as effective platforms to exchange thoughts on the latest technical or regulatory developments in pathology, discuss interesting cases and images with colleagues, and communicate with patients in dedicated support groups. The number of the pathologists engaging in social media - let’s call them ‘social network pathologists’ - continues to grow swiftly. On Twitter alone, there were over 19,000 Tweets on #Pathology in the month of September 20171, leading influencers in the field have thousands of followers and even a formal ontology on Twitter hashtags for pathology subspecialties 2emerged.
Looking around the web, we find numerous practical how-to guides3 for the use of social media as medical doctor and pathologist, including tips of how to prevent pitfalls to HIPAA’s regulations4. But what are the real drivers for pathologists to publicly share, present and discuss their work and to mingle in these virtual groups?
Is it the need for more expert discussion and exchange, which healthcare IT solutions cannot satisfy? Is it the community spirit or a self-branding and advocating-opportunity for one’s institution or personal career? Or is it a mix of all these reasons? Let’s try to get to the bottom of the most apparent benefits of social media for pathologists and understand what motivates them to ‘read, like and tweet’.
SOCIAL NETWORK PATHOLOGIST DRIVER #1: CLOSING THE COMMUNICATION GAP
Social media provides us with simple means to communicate and share viewpoints, ideas and experiences with others - anywhere, instantly and at zero costs. It is therefore not at all surprising, that a field of medicine, that embraces the interpretation of images at its core, jumped on this channel to close the communication gap between stakeholders inside and outside their lab or hospital. Software providers in the field of digital pathology focus on work-flow solutions that are dedicated to collecting Protected Health Information. Public availability, which is a must to foster inter-institutional exchange, is mainly forbidden by design6. As a result, interaction with treating physicians, specialists and colleagues working in other institutes and even with patients shifted to tools that are available to everybody.
By using social media apps such as Facebook and Twitter, pathologists can instantly and efficiently share their case images (encoded/de-identified for privacy), give and ask for opinions on photomicrographs, and comment on others’ posts - anytime, anywhere, from any device and even outside of their daily routine work after hours. Usage is free of charge, no budget decisions need to be taken, no IT department to be involved.
While the interaction between pathologists on social media usually also includes exchange on new regulations, medical trends, educational videos and content links, the focus seems to rest on sharing and discussing interesting, rare and ambiguous histology images with experts around the world. Although annotations to posted histology images, replies to questions and content shared on social media can and will never be basis to a diagnosis, pathologist widely welcome the value of such comments for supporting and confirming the diagnosis of human disease.
Pathologists' drivers to use social media - a summary. Copyright: microDimensions
Not only for regions, where pathologist expertise is widely scattered, exchanging additional opinions about a case can make a difference to the quality of patient treatment. With the urge to make diagnosis more precise, especially general pathologists consult with colleagues qualified in subspecialties. However, direct communication with their personal network is cumbersome and subject to both technical and availability constraints. By posting a snapshot taken with their smartphone from the microscope or even a screenshot of a dedicated area of the whole slide image on Twitter, pathologists can open up the possibility to receive additional opinions from virtually all experts of the field worldwide at a glance.
Social network pathologist driver #2: Explore and build pathological knowledge
Digitizing medical and pathological knowledge and thereby giving broad access as a resource can only be an advantage: Building online libraries of medical publications was one step on the digital journey. An additional potential to explore and build pathological knowledge unfolds in what is aggregated and made publicly available on social media: images with comments and annotations, rare cases and photomicrographs suitable for training. Especially in developing countries, or in fact anywhere where educational resources are scarce, and books are too expensive, pathologists value the knowledge available free of charge and instantly via social media channel.
Residents join social groups or follow long-standing experts (‘influencers of subspecialty’) to expand their knowledge, use it as educational training field, learn and absorb. At the same time, pathologists constantly come across more complex or rare cases outside their particular expertise. Leveraging the extensive public networks, they can get easy and fast confirmation of their initial assessment, or at least be guided into a previously unseen direction.
SOCIAL NETWORK PATHOLOGIST DRIVER #3: BEING PART OF THE COMMUNITY!
Engaging in social media networks and groups makes us feel part of a community. Whenever pathologists collaborate with experts of the same profession via social media, they contribute or take benefit from the community knowledge. This fosters a unique feeling of belonging and playing a role in the huge complex healthcare system. One of the ultimate goals of social networks is, to extend the own professional network, eventually turning an initial virtual ‘like’, ‘follow’ or ‘reply’ into a real physical, beneficial interaction. Just lately, I have overheard a conversation between two young pathologists who delightfully met for the first time at the European Congress of Pathology in Amsterdam after starting and intensifying their Facebook friendship. Think of the opportunities that open up: joint publication projects, long-term second-opinion collaboration, or international career opportunities.
Live tweets on pathology events and conferences are a welcome form of exchange - not only to learn what went on in a parallel conference session. They are not only fostering the community of participants, voice opinions about the event and keep memories alive. Social media is also used to continue discussion initiated in sessions or poster presentations, letting the entire pathology community join in. And in fact, the United States and Canadian Academy of Pathology (USCAP) 2015 annual meeting gathered over 660 tweeting participants resulting in more than impressive 6,500 live tweets about the event6.
SOCIAL NETWORK PATHOLOGIST DRIVER #4: A TENDENCY TO CLUSTER AND THE MERITS OF GROUP COLLABORATION
More and more of the group collaboration is turning to social media, in particular to Facebook groups, as they “provided a no-cost way for pathologists and others across the world to interact online with many colleagues”7. Here, various topics including histology findings can be discussed in a closed, hence locally unlimited real-time setting.
The objectives to form, join and contribute to a group are manifold. Some groups form around specific subspecialties, giving space to focused exchange in that field. There are groups of regional and international organizations and associations that can attract over ten thousands of members. Then, there are smaller, often temporary groups that are means of communication during the preparation of papers or publications. In education, residents can turn to groups to ask their study questions or discuss their educational success online with their fellow students from other universities.
At this point I would also like to mention the smallest group format on social media: private chat possibilities, which most social media channels provide. Tools such as private message (PM) in Facebook or Direct Message in Twitter enable pathologists to continue public or group-wide discussions in a very private environment. It is common, that a post turns into a lively one-to-one discussion between the posting pathologist and a commenter - or even between commenters. This hints us to the next driver for pathologists using social media: building the own professional network and career.
Social network pathologist driver #5: Self-branding, institute-advocacy and other career kicks
As a private person we engage in social media to inform our networks about what we are currently doing and especially what we are doing well. The social channels’ power to present and self-market ourselves and our employers is priceless. Every post a pathologist makes has the potential to identify him or her as recognized expert in a subfield and suitable to give a qualified second opinion, eligible to teaching opportunities or candidate to give speeches - turning a contribution to the healthcare community into career kicks for the social network pathologist.
Unlike large hospitals, not many private pathological institutes have dedicated manpower to run marketing initiatives for their organizations. When it comes to international reputation building, communication of the latest research results or simply promoting vacancies, the social media engagement of their pathology staff can be a real asset and a win-win situation for both employee and employer.
As a positive side effect, communicating about pathology work publicly contributes to the image of the pathologist profession as such, its role for a more precise and profound health care and it also helps to break down stereotypes - check out #ILookLikeAPathologist on Twitter.
Summary
No social network pathologist’s motivation is driven by only one of the above named goals. Usually their interaction on social channels combines all of them and even mixes with private interest posts and tweets. What unifies them - however - couldn’t be a more noble intention: to understand disease and help people to heal people.
1 real-time analytics data for past 30 days on 29 September 2017: https://www.symplur.com/healthcare-hashtags/ontology/pathology/
2 https://www.symplur.com/healthcare-hashtags/ontology/pathology/
3 E.g. Jerad M Gardner, MD: http://pathinfo.wikia.com/wiki/Social_Media_Guide_for_Physicians, http://pathinfo.wikia.com/wiki/Social_Media_Guide_for_Pathologists
4 E.g.: http://blog.intakeq.com/5-ways-to-stay-hipaa-compliant-when-using-social-media, May 2017
5 As an alternative, all stakeholders would need to work with the same DP system, which is not very realistic; and standardized exchange protocols are not yet established.
6 Modern Pathology (2017): #InSituPathologists: how the #USCAP2015 meeting went viral on Twitter and founded the social media movement for the United States and Canadian Academy of Pathology, http://www.nature.com/modpathol/journal/v30/n2/full/modpathol2016223a.html
7 Gonzalez et al: Facebook Discussion Groups Provide a Robust Worldwide Platform for Free Pathology Education, Arch Pathol Lab Med 141, pp. 690-695, May 2017
Further Reading on Social Media in Pathology
- If You Are Not on Social Media, Here’s What You’re Missing! #DoTheThing http://archivesofpathology.org/doi/pdf/10.5858/arpa.2016-0612-SA?code=coap-site
- Social Media Guide for Pathologists: http://pathinfo.wikia.com/wiki/Social_Media_Guide_for_Pathologists
- Pathology Image-Sharing on Social Media: Recommendations for Protecting Privacy While Motivating Education: http://journalofethics.ama-assn.org/2016/08/stas1-1608.html
- #InSituPathologists: how the #USCAP2015 meeting went viral on Twitter and founded the social media movement for the United States and Canadian Academy of Pathology: http://www.nature.com/modpathol/journal/v30/n2/full/modpathol2016223a.html
- Fine social aspiration: Twitter as a voice for cytopathology: http://onlinelibrary.wiley.com/doi/10.1002/dc.23713/epdf